Computational methods for efficiently simulating ecological communities are increasingly in demand. In 2008, James Rosindell popularised coalescence methods for simulating neutral ecological communities. These methods can be orders of magnitude faster than the alternatives, and can make spatially explicit neutral simulations on an effectively infinite landscape feasible. However, the technical challenge of implementing coalescence methods has left them out of reach of many ecologists. Now, Sam Thompson, who recently graduated from the lab with a PhD, being co-supervised by Rosindell at Imperial College London, has published user-friendly high-performance software packages for performing coalescence simulations in R and Python. The packages are described in a new paper in Methods in Ecology and Evolution.
The essence of coalescence methods is that they run backwards in time, which increases efficiency because only past lineages that lead directly to individuals within the focal community in the present day need to be tracked. In contrast, with standard forward-time-simulations, a much larger number of lineages needs to be tracked because it is not known in advance which ones will persist and be part of the focal community in the future. The catch is that coalescence can only be applied to particular types of ecological models, in particular to neutral models where an individual’s chances of birth and death are independent of its species identity. Coalescence techniques were originally invented in population genetics, and then imported to ecology by Rosindell.
Sam’s new software packages include the basic features from Rosindell’s 2008 paper, along with many enhancements and novel features, including varying individual density across a landscape, changing landscape configuration over time, and a variety of dispersal and speciation modes. The software was also used in Sam’s recent Ecology Letters paper on extinction debt. The core package code is written in C++, but the R and Python packages give a user-friendly interface, that should enable easy uptake by any ecologists interested in running fast spatially explicit neutral simulations.
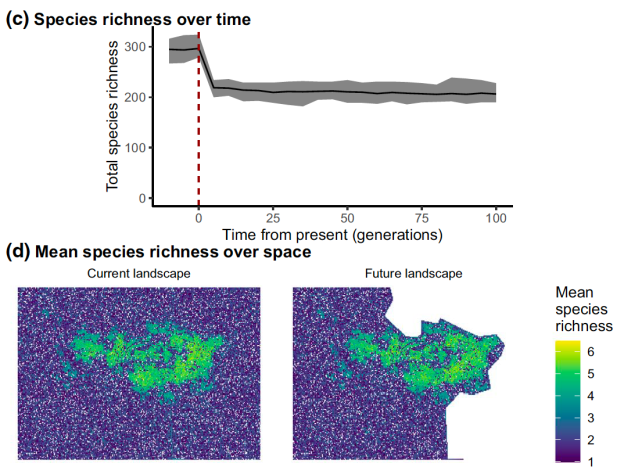
Output from the new software packages, showing (c) change in species richness over time in a hypothetical scenario after (d) part of a landscape surrounding a national park (green area) has been cleared.
